Projects
Intelligent Cell Segmentation for Spatial Technologies
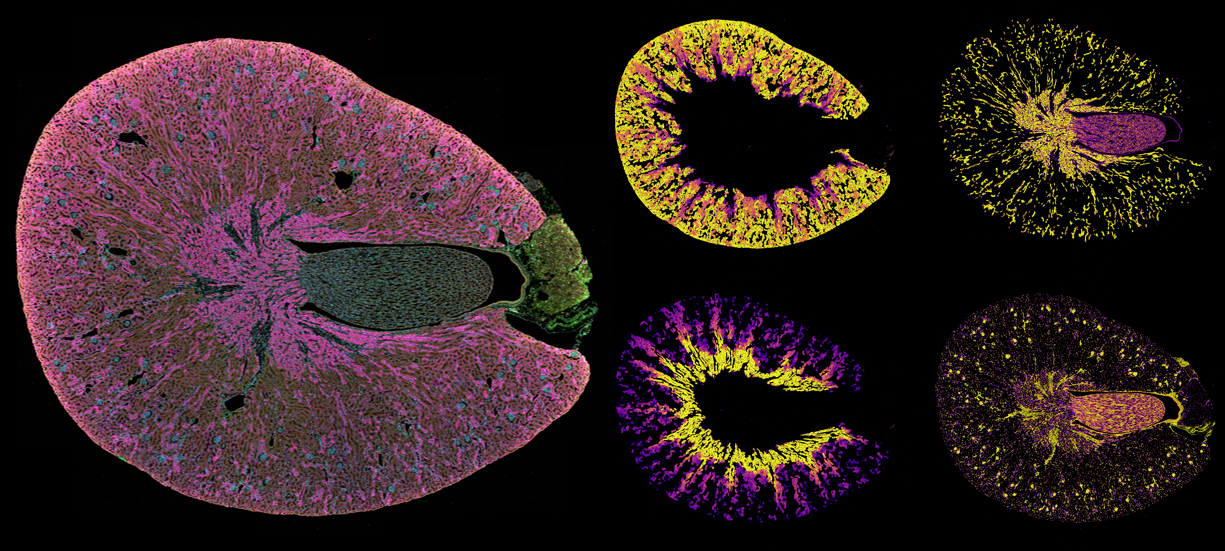
Precise cellular segmentation is foundational to every downstream spatial analysis. At AIMS@USC, we are advancing next-generation segmentation frameworks that leverage machine learning and computer vision to move beyond rule-based approaches. By integrating probabilistic modeling, deep learning architectures, and multimodal image features, we aim to improve accuracy, scalability, and biological fidelity across diverse tissue contexts. Our vision is to develop adaptable, platform-agnostic segmentation pipelines that not only delineate cells more accurately, but also quantify uncertainty—transforming segmentation from a preprocessing step into a scientifically rigorous, data-informed component of spatial discovery.
Cross-Dataset Label Transfer and Cell Type Harmonization
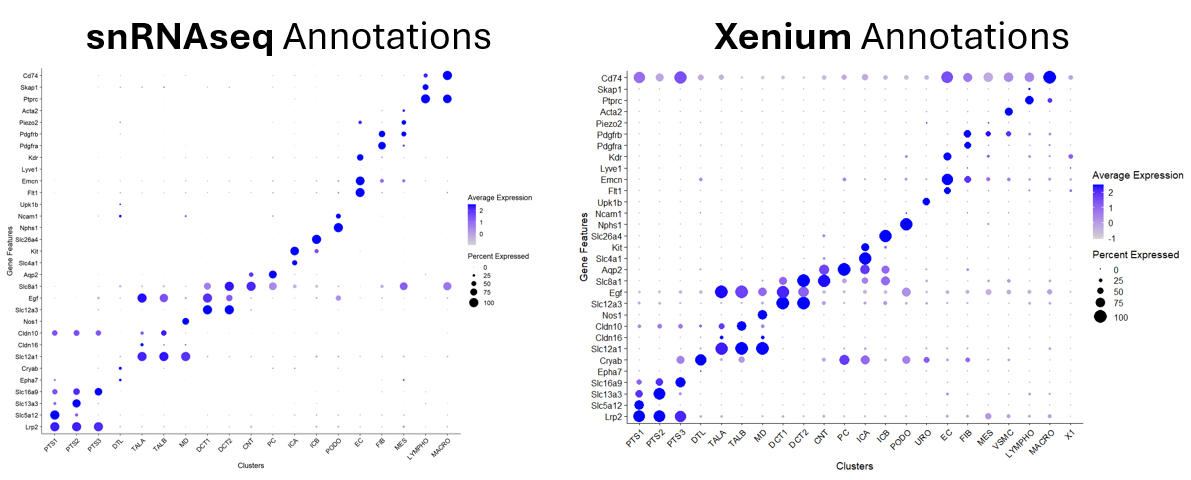
As spatial and single-cell datasets proliferate, the ability to accurately transfer cell type annotations across platforms has become essential. AIMS@USC is building robust computational frameworks for label transfer that integrate transcriptomic similarity, spatial context, and domain adaptation techniques. Our goal is to move beyond simple nearest-neighbor mapping toward biologically informed, uncertainty-aware annotation systems. By harmonizing datasets across technologies, cohorts, and laboratories, we aim to enable reproducible cell type definitions and accelerate integrative discovery across studies, diseases, and populations.
Integrating Dissociated and Spatial Transcriptomic Data
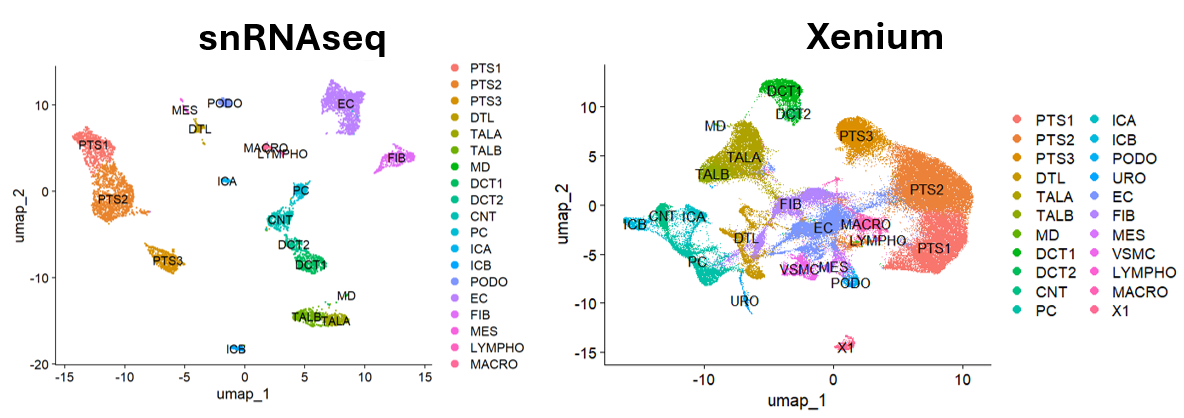
Dissociated single-cell sequencing offers depth, while spatial technologies preserve context. At AIMS@USC, we seek to bridge these complementary modalities through principled statistical modeling and AI-driven integration strategies. By systematically comparing dissociated and spatial datasets, we aim to identify modality-specific biases, quantify information loss, and develop correction frameworks that unify molecular resolution with tissue architecture. Our long-term vision is to create integrative maps that reconcile cellular identity, state, and microenvironment—providing a more complete and mechanistic understanding of complex biological systems.